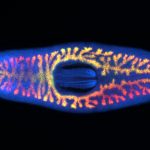
David Forsthoefel, Ph.D.
Assistant Professor
Genes & Human Disease Research Program
My 101
Injury, aging and disease can damage internal organs and other body parts, but our bodies often fail when trying to repair wounded or worn-out tissues. By contrast, many animals like hydra, starfish, flatworms, zebrafish, lampreys and salamanders have the amazing capacity to repair or even replace damaged tissues. In the last 20 years, innovative technologies have been developed that enable us to study the roles of thousands of genes at a time. These technologies, known as “functional genomics,” have given us an incredible new window into the mysterious secrets of animal regeneration. Because regenerating animals and humans have thousands of similar genes in their DNA, what we learn from these organisms will tell us much more about how to build, repair and maintain healthy human tissues and organs.
The Forsthoefel Lab uses functional genomic approaches to understand regeneration in freshwater flatworms called planarians. Planarians have astonishing regenerative abilities that have fascinated biologists for centuries. After amputation, they can grow new heads and tails (and all of their internal organs) effortlessly, in a process that takes about a week. You can even cut a planarian into many tiny pieces, and each piece will regenerate into a perfectly normal planarian. One of the keys to planarian regeneration is that they possess adult stem cells, found throughout their bodies, that sense injury and get to work rebuilding organs every time they are damaged.
Our research focuses on the planarian intestine. How do adult stem cells know when the intestine is injured? How are just the right number of new intestinal cells made, and how do they assemble into a functional digestive tract? How does the intestine communicate with other tissues, in order to signal healthy or injured status? Answers to these and related questions could one day help us to improve treatments for human digestive system diseases. For example, do planarians control cell division in unique ways that might tell us more about colon cancer? How do planarians cope with stresses that cause Crohn’s disease and ulcerative colitis? By discovering the fundamental rules of regeneration, and by studying genetic and biochemical processes implicated in human disease, we hope to learn some of the secrets these amazing little flatworms have to tell us.
Research
Many animals continuously renew the epithelial linings of their digestive tracts in response to normal cellular turnover, or “wear and tear”. However, only some -- like salamanders, sea cucumbers and flatworms -- can repair more severe injury to gastrointestinal (GI) organs, such as transection or tissue loss. Although our understanding of homeostatic renewal has advanced considerably, regeneration of digestive organs remains poorly characterized. Investigations of GI regeneration will therefore broaden our understanding of fundamental, universal mechanisms of tissue repair. In addition, such studies are relevant in specific medical contexts, such as short bowel syndrome, in which surgical removal of portions of a patient’s intestine leads to impaired nutrient absorption. Furthermore, studies of GI regeneration will generate new insights into proliferation, differentiation and other cellular processes whose dysregulation underlies common human pathologies, including inflammatory bowel disease, Barrett's esophagus and colorectal cancer.
The Forsthoefel Lab exploits the gastrovascular system, or intestine, of the planarian Schmidtea mediterranea to elucidate fundamental and biomedically relevant mechanisms of regeneration. Planarians are free-living, freshwater flatworms with astonishing regenerative abilities: even small tissue fragments completely regenerate into viable animals with functional internal organs. In planarians, regeneration is achieved through the activation of pluripotent somatic (adult) stem cells called neoblasts, as well as dramatic remodeling of pre-existing tissue. In recent years, the development of molecular tools and functional genomic resources have made S. mediterranea a robust animal model in which to understand stem cell-driven regeneration. Our research focuses on several key areas:
Microenvironment regulation during regeneration. Cell non-autonomous signaling influences cellular proliferation and differentiation, particularly in the local environments surrounding adult stem cells, called niches. How niches are modulated in response to tissue damage, however, is not well understood. Our previous work (Forsthoefel et al., Dev. Cell, 2012) suggests that the planarian intestine plays an important role in coordinating neoblast dynamics. We are coupling cell sorting and laser capture microdissection with functional genomics techniques in order to identify signals that influence stem cell dynamics, and to elucidate the transcriptional programs that control their expression.
Mechanisms of stem cell-driven organ regeneration. Understanding organ regeneration requires elucidation of the genetic programs that generate appropriate numbers of distinct cell types in a post-embryonic context. How do stem cells and intestinal progenitors respond to tissue damage? How are evolutionarily conserved mechanisms used to specify secretory and absorptive lineages during regeneration of digestive organs? Do regenerating animals regulate proliferation and differentiation in unique ways in response to injury? We are using a variety of approaches to address these questions.
Modeling gastrointestinal disease in planarians. Little is known about how genes implicated in human gastrointestinal diseases function during regeneration. How do planarians control the rate of stem cell division so that new intestinal cells are produced in appropriate numbers, without hyperplasia? How does the planarian digestive epithelium respond to stresses that cause human diseases such as Crohn’s and ulcerative colitis? Our goal is to apply functional genomics in novel ways to develop deeper, medically relevant insights into the pathogenesis of colon cancer and inflammatory bowel disease.
Brief CV
Education
B.S., Xavier University, Cincinnati, Ohio, 1994
Ph.D., The Ohio State University, Columbus, Ohio, 2005
Postdoc, University of Illinois at Urbana-Champaign and Howard Hughes Medical Institute, 2005-2016
Honors and Awards
cum laude and University Scholar, Xavier University, 1994
Best talk, Falkenthal Graduate Research Symposium, The Ohio State University, 2003
Herta Camerer Gross Graduate Summer Research Fellowship, The Ohio State University, 2004
Best postdoctoral talk, Midwest Society for Developmental Biology Meeting, 2008
Travel award, Society for Developmental Biology Meeting, 2008
NIH Ruth L. Kirschstein Postdoctoral National Research Service Award, NIDDK, 2006-2009
Other Activities
Contributor, Project NEURON, University of Illinois at Urbana-Champaign, 2010
Mentor, Summer Research Opportunities Program, University of Illinois at Urbana-Champaign, 2014
Faculty Instructor (invited), Frontiers in Stem Cells and Regeneration Advanced Training Course, Marine Biological Laboratory, Woods Hole, Massachusetts, 2014-2022
Faculty Instructor (invited), Frontiers in Aging and Regeneration Research Advanced Training Course; Morehouse School of Medicine, Atlanta, 2017; Xavier University, New Orleans, 2018
Faculty Mentor, OMRF Fleming Scholar Program, 2019; 2021
Faculty Mentor, Oklahoma School for Science and Math Research Mentorship Program, 2018-2022
Faculty Member, OMRF Postdoc Training Committee, 2018-2022
Faculty Member, Graduate Education Committee, Dept. of Cell Biology, University of Oklahoma Health Sciences Center, 2022
Membership
Society for Developmental Biology
American Society for Cell Biology
International Society for Extracellular Vesicles
Joined OMRF scientific staff in 2016
Publications
Recent Publications
Wong LL, Bruxvoort CG, Cejda NI, Delaney MR, Otero JR, Forsthoefel DJ. Intestine-enriched apolipoprotein b orthologs are required for stem cell progeny differentiation and regeneration in planarians. Nat Commun 13:3803, 2022 July, PMID: 35778403, PMCID: PMC9249923
Avalos PN, Forsthoefel DJ. An Emerging Frontier in Intercellular Communication: Extracellular Vesicles in Regeneration. Front Cell Dev Biol 10:849905, 2022 May, PMID: 35646926, PMCID: PMC9130466
Forsthoefel DJ, Cejda NI, Khan UW, Newmark PA. Cell-type diversity and regionalized gene expression in the planarian intestine. Elife 9, 2020 April, PMID: 32240093, PMCID: PMC7117911
Selected Publications
Forsthoefel DJ, Waters FA, Newmark PA. Generation of cell type-specific monoclonal antibodies for the planarian and optimization of sample processing for immunolabeling. BMC Dev Biol 14:45, 2014 Dec, PMID: 25528559, PMCID: PMC4299570.
Forsthoefel DJ, James NP, Escobar DJ, Stary JM, Vieira AP, Waters FA, Newmark PA. An RNAi screen reveals intestinal regulators of branching morphogenesis, differentiation, and stem cell proliferation in planarians. Dev Cell. 2012 Oct 16;23(4):691-704. PMID: 23079596; PMCID: PMC3521571.
Forsthoefel DJ, Park AE, Newmark PA. Stem cell-based growth, regeneration, and remodeling of the planarian intestine. Dev Biol. 2011 Aug 15;356(2):445-59. PMID: 21664348 PMCID: PMC3490491
Forsthoefel DJ, Newmark PA. Emerging patterns in planarian regeneration. Curr Opin Genet Dev. 2009 Aug;19(4):412-20. PMID: 19574035 PMCID: PMC2882238
Forsthoefel DJ, Liebl EC, Kolodziej PA, Seeger MA. The Abelson tyrosine kinase, the Trio GEF and Enabled interact with the Netrin receptor Frazzled in Drosophila. Development. 2005 Apr;132(8):1983-94. PMID: 15790972
Liebl EC, Forsthoefel DJ, Franco LS, Sample SH, Hess JE, Cowger JA, Chandler MP, Shupert AM, Seeger MA. Dosage-sensitive, reciprocal genetic interactions between the Abl tyrosine kinase and the putative GEF trio reveal trio's role in axon pathfinding. Neuron. 2000 Apr;26(1):107-18. PMID: 10798396
Contact
Genes & Human Disease, MS 42
Oklahoma Medical Research Foundation
825 N.E. 13th Street
Oklahoma City, OK 73104
Contact Information
Phone: (405) 271-4047
Fax: (405) 271-3765
E-mail: David-Forsthoefel@omrf.org
For media inquiries, please contact OMRF’s Office of Public Affairs at news@omrf.org.
Lab Staff
Lily Wong, Ph.D.
Staff Scientist
Matthew Delaney
Research Technician II
Priscilla Avalos
Graduate Student
Susan "Suzy" Collins
Project Coordinator II
News from the Forsthoefel lab

July A pair of OMRF scientists take a different approach to the study of healing. Time, it’s said, heals all wounds. But we all know that’s not quite true. “Everything must work just right for proper wound healing to take place,” says OMRF’s Dr. Lorin Olson. “It’s a delicate balance.” Olson studies a compound in the […]

So, 2020. Not quite the 366 days any of us were expecting, huh? But we all found a way to muddle through. And the scientists, physicians and staff of OMRF were no different. For most employees, the emergence of Covid-19 in March brought a months-long shutdown of onsite activities. Throughout the spring, only a handful […]