Expanding Excellence in Developmental Biology in Oklahoma
Welcome to our website. We have completed Phase I and are currently in the final year of Phase II of our COBRE: “Expanding Excellence in Developmental Biology in Oklahoma”. Developmental Biology is fundamental to progress in determining the molecular basis of many diseases, as well as for the rapidly emerging field of regenerative medicine. Discoveries made in the field of Developmental Biology will enable new therapeutic strategies for a myriad of human diseases. The goal of our COBRE has been to recruit and train promising Project Leaders to build a critical mass of Developmental Biologist in Oklahoma. We succeeded in building an interactive community where developmental biologists can flourish and have established core facilities to provide critical tools for making fundamental discoveries that can be translated into technologies benefiting human health.
In Phase I we recruited five scientists using a variety of model systems to zebrafish, Drosophila, mice and adult stem cells. All five of our Phase I Junior Investigators now run well-funded research laboratories at the Oklahoma Medical Research Foundation (OMRF) or the University of Oklahoma Health Sciences Center (OUHSC).
In Phase II we recruited five new Junior Investigators, four of these have graduated. We recruited two additional scientists and currently support three Phase II Project Leaders. Our Phase II Project Leaders have done outstanding work addressing fundamental questions in developmental biology (cell growth and division, cell signaling, differentiation, tissue regeneration, and modeling human diseases in model organisms) with translational relevance, using a variety of model systems (planarians, Drosophila, C. elegans, zebrafish, mice and cell culture systems). This infusion of new experimental strategies and zeal has greatly benefited our institution overall.
In addition to building our infrastructure, Phase II COBRE support is addressing important and alarming nation-wide trends in the biomedical research workforce: the increasing age of investigators receiving initial NIH support (current average of 42 years), a high dropout rate for those awardees (up to 50%) and difficulty in remaining competitive over time. Erosion of NIH support for investigator-initiated proposals in favor of program awards and expansions in the requirements for publications in scientific journals require more rapid affiliation with large teams. However, that can conflict with the need to achieve independent recognition. Retaining rising stars in an increasingly competitive environment represents an additional challenge for institutions. Our strong track record of successful research and high investigator retention indicate that OMRF can bring unique resources to address these challenges while growing an exciting, multi-disciplinary research focus in Developmental Biology. In Phase II, this has been accomplished through the following Specific Aims:
- Aim 1. To grow our multi-disciplinary research program in Developmental Biology through the nurturing of a talented group of Junior Investigators with outstanding potential to become leaders in their fields.
- Aim 2. To mentor the Junior Investigators, using a combination of senior scientists and recently graduated COBRE Junior Investigators within existing OMRF and University of Oklahoma faculty.
- Aim 3. To form an External Advisory Committee of internationally recognized developmental biologists to evaluate the progress of, and offer advice to, the Junior Investigators.
- Aim 4. To maintain and expand Core facilities to support the research of the Junior Investigators and lay the groundwork for sustainable operation after COBRE funding ends.
- Aim 5. To promote scientific interactions among the Junior Investigators and create an environment conducive to investigator retention.



Administrative Core
The goal of the Administrative Core is to coordinate the scientific cores. To accomplish this goal, the Administrative core will select, fund, and monitor pilot projects, maintain the mentoring environment, and provide logistical support for research.
External Advisory Committee
Roger Markwald, Ph.D., Chair
Carla Kim, Ph.D.
Leonidas Tsiokas, Ph.D.
Current Projects
Notch regulation of COPD-associated goblet cell hyperplasia/metaplasia
Matthew Walters Ph.D.
Associate Professor of Medicine, OUHC
Chronic obstructive pulmonary disease (COPD) is the 3rd leading cause of death in the US with >90% of cases caused by cigarette smoking. Currently there is no cure for COPD. Remodeling of airway epithelial architecture in response to cigarette smoke exposure plays a critical role in the pathogenesis of COPD. Goblet cell hyperplasia or metaplasia is an epithelial remodeling phenotype characterized by increased numbers of mucus-producing (“goblet”) secretory cells. In association with decreased ciliated cell numbers, these changes culminate in excess mucus production and impaired mucociliary clearance, which contribute significantly to COPD morbidity and mortality. Therefore, identifying the mechanisms by which cigarette smoke dysregulates secretory cell differentiation in the airway epithelium is critical to developing novel therapeutics to treat COPD.
Basal cells (BC) function as tissue-specific stem/progenitor cells in the human airway epithelium. Notch signaling is a developmental pathway that plays a key role in regulating BC stem/progenitor function in both humans and mice. This includes regulating the balance of BC differentiation into either secretory or ciliated cells. In vivo human studies have identified alterations in Notch signaling components at the mRNA, protein and epigenetic levels in the small airway epithelium (SAE) of smokers with and without COPD relative to nonsmokers. These include increased levels of activated NOTCH1 receptor and altered expression of NOTCH3 and multiple Notch downstream effector genes (e.g., HES5, HEY1 and HEY2). We demonstrated in vitro that constitutive activation via the NOTCH1 and 3 receptors in nonsmoker-BC activates a unique transcriptional response relative to NOTCH2 and 4, with increased expression of the downstream effectors HES5, HEY1, HEY2 and HEYL. Furthermore, NOTCH1 and 3 signaling activation increased secretory cell differentiation with a corresponding decrease in ciliated cell differentiation characteristic of goblet cell hyperplasia/metaplasia. Therefore, we hypothesize that cigarette smoking increases Notch signaling activity in BC stem/progenitors resulting in development of goblet cell hyperplasia/metaplasia in the airways of smokers with COPD.
Our long-term goal is to uncover pathogenic mechanisms and treatment strategies for COPD. Although Notch regulates BC differentiation and is implicated in COPD, there is limited insight into how Notch signaling contributes to COPD. Therefore, despite Notch providing a potential therapeutic target to treat COPD-associated goblet cell hyperplasia/metaplasia, important questions remain. First, what is the effect of cigarette smoking on Notch activity in the airway epithelium (Aim 1)? Second, as BC from COPD patients have altered regenerative capacity compared to BC from nonsmokers, does modulation of Notch activity in COPD-BC have the same outcome as in nonsmoker-BC (Aim 2)? Third, what are the downstream effectors that regulate Notch-mediated goblet cell differentiation (Aim 3)? To address these questions we propose the following Aims:
1) Determine whether cigarette smoke exposure activates Notch signaling in the airway epithelium in vitro and in vivo.
2) Determine whether BC stem/progenitor cells from nonsmokers vs. patients with COPD respond differentially to Notch signaling modulation as they differentiate into a mucociliated epithelium.
3) Determine whether specific Notch downstream effectors can induce goblet cell differentiation in vitro and whether targeting these genes can inhibit Notch-induced goblet cell hyperplasia/metaplasia.
The outcomes of these studies will advance our understanding of the impact of cigarette smoke exposure on Notch activity in the airway epithelium and identify Notch effectors that regulate goblet cell differentiation. Further, they will provide candidate therapeutic targets to suppress COPD-associated goblet cell hyperplasia/metaplasia.
Mechanistic insights into how the pBAF complex influences chromatin structure
Jake Kirkland, Ph.D.
Assistant Member, Cell Cycle & Cancer Biology, OMRF
Within the nucleus, DNA is packaged into chromatin states that vary in accessibility. Importantly, accessibility allows DNA binding proteins such as transcription factors, and repair and replication proteins to contact the DNA fiber. Chromatin regulators set up, maintain, and reverse chromatin states thereby controlling gene expression, chromatin structure, and genome fidelity. Understanding the mechanisms by which chromatin states are modulated is medically significant because approximately 50% of tumors have alterations in chromatin regulators. The family of chromatin regulators that is the focus of this proposal, the mammalian SWI/SNF family (also known as BAF complexes), is mutated in 20% of all tumors. BAF complexes are combinatorially assembled complexes containing 12-15 subunits encoded by 29 genes. In mammals there are 3 distinct major forms: canonical BAF (cBAF), Polybromo-associated BAF (pBAF), and GLTSCR-associated BAF (gBAF). All three share common subunits but also contain unique subunits. Of the three major BAF subtypes, the cBAF complex has the best understood role in chromatin regulation. However, much less is known about the mechanisms and function of the pBAF complex, including how common cancer mutations or subunit deletions affect the complex. A multitude of cancer genomics studies shows that alterations in cBAF and pBAF are frequent, but largely occur in cancers of different tissue origins suggesting divergent roles. On a genome-wide level cBAF most often acts at promoters and enhancers to open up chromatin leading to a transcriptionally competent locus. While pBAF is found at enhancers and promoters, it is also found along gene bodies. pBAF localization correlates with both transcriptional activation and to a larger extent than cBAF to transcriptional repression. In contrast to cBAF, pBAF may work cooperatively with polycomb during DNA repair, however the generality of this in other cell processes remains unknown.
Polycomb proteins act to oppose transcription, through histone modifications, recruitment of other proteins, and local chromatin organization changes. While the cBAF complex has many different roles in the cell, recent studies of cBAF mechanism have shown that one of the primary roles is to oppose polycomb proteins, thereby creating a more accessible locus. In most cases accessibility alone is not sufficient for transcriptional activation. However, I found that cBAF recruitment using a Cas9 based chemical induced proximity approach (FIRE-Cas9) is sufficient for transcriptional activation of the mouse Nkx2-9 locus. I can now use Nkx2-9 as a model locus of chromatin and transcription changes just minutes after inducible SWI/SNF recruitment. While it is known that cBAF and/or gBAF complexes opposes polycomb when recruited to Nkx2-9, what is not known is how recruitment of pBAF affects polycomb, the chromatin state in general or the transcriptional state of Nkx2-9.
My long-term goal is to define the mechanistic differences between the mSWI/SNF pBAF and cBAF/gBAF complexes in regard to polycomb localization, histone modifications, chromatin accessibility, transcription, and genome stability. My overall objective is to understand how the similar but distinct BAF complexes act through unique mechanisms mediated by specific subunits. Knockout of different individual subunits in mSWI/SNF complexes leads to disparate transcriptional and polycomb changes across the genome. This observation led to my central hypothesis that oncogenic alterations in subunits of similar complexes contribute to disease through distinct mechanisms. The rationale for the proposed research is that once we know mechanistic differences between similar complexes, we will have a better understanding of oncogenic mechanisms and be able to better predict points of pharmacological attack in order to restore a more normal balance of chromatin states.
I plan to test my central hypothesis by pursuing the following two specific aims:
1) Characterize genomic alterations after acute loss of the pBAF complex.
2) Compare the kinetics and mechanisms of cBAF vs. pBAF mediated chromatin changes.
With respect to outcomes, I expect the differences in mechanism between these similar, but divergent complexes will lead to a better understanding of alterations in chromatin structure that lead to an oncogenic state. This will have a positive translational impact because a better understanding of the relationship between chromatin regulators, cancer subtypes, and chemotherapeutic regimes will be important for better patient responses and outcomes. This Award will provide me the resources and training necessary to compete independently for NIH individual research grants and other external peer-reviewed support.
Determining the mechanistic effects of the sas-6(L69T) mutation in C. elegans
Jyoti Iyer, Ph.D.
Assistant Professor of Chemistry & Biochemistry, University of Tulsa
Mutations in centriole duplication (CD) genes are associated with diseases such as primary microcephaly (MCPH), and cancer. To aid in the development of therapy against these diseases, it is critical to understand how CD proteins normally function within cells. Studying the effects of disease-associated mutations in CD genes can provide important clues into their biological roles. One such mutation in the sas-6 gene (HsSAS6(I62T)) was associated with the incidence of MCPH in a Pakistani family. SAS-6 function is essential for both primary ciliogenesis and CD: defects in each of these processes have been implicated to cause MCPH. Although the MCPH phenotype caused by the HsSAS6(I62T) allele indicates a potential defect in primary cilium function and/or CD, the mechanistic failures caused by this mutation are unknown. Importantly, the effects of this MCPH-associated mutation on ciliogenesis have never been investigated. Over-expression of the HsSAS6(I62T) mutation in U2OS cells yielded monopolar spindles. This suggests that when over-expressed, the mutant protein can have a dominant negative effect on CD through an as yet unknown mechanism. The molecular and developmental consequences of this mutation have never been investigated in the context of a whole multicellular eukaryotic model organism. The function of the SAS-6 protein is conserved among several different species, including humans and C. elegans. In fact, SAS-6 was first identified in C. elegans as being essential for proper CD and was later shown to have a similar role in humans. SAS-6 function is also essential for ciliogenesis in both C. elegans and humans. Although HsSAS-6/SAS-6 is widely recognized as a critical structural component of the centrioles, many gaps still exist in our understanding of how SAS-6 mediates primary ciliogenesis and CD in vivo. One reason for this is that since sas-6 is an essential gene, severely impairing sas-6 activity causes embryonic lethality. Our lab has generated a C. elegans strain carrying the human MCPH-equivalent sas-6(L69T) mutation using CRISPR/Cas9 genome editing. Our preliminary data indicate that this mutation mildly perturbs CD in C. elegans as appears to be the case in humans. By generating a sas-6 allele that has a pathological relevance, we have created an excellent tool to probe the function of SAS-6 in the processes of ciliogenesis and CD. Therefore, in this proposal, we will utilize this new, health-relevant sas-6 allele to study the role of SAS-6 in ciliogenesis and CD.
The long-term goal of my laboratory is to study the effect of disease-associated mutations in conserved ciliogenesis and CD genes. The objectives in this endeavor are to determine the effects of the sas-6(L69T) mutation on C. elegans ciliogenesis and CD. Our preliminary data demonstrate that, the presence of the sas-6(L69T) mutation increases CD failure of a strain that is partially compromised for CD. Based upon these and other preliminary data, our central hypothesis is that the sas-6(L69T) mutation inhibits SAS-6 function leading to impaired ciliogenesis and CD. The rationale for this proposal is that since the human equivalent of the sas-6(L69T) mutation has a pathological consequence, this mutation will provide important insights into the normal biological function of SAS-6. To test our central hypothesis, we will pursue the following Specific Aims:
1) Determine the effect of the sas-6(L69T) mutation on C. elegans ciliogenesis.
2) Elucidate the molecular mechanism by which the sas-6(L69T) mutation impairs CD in C. elegans.
Upon the successful completion of these aims, the expected outcomes are an enhanced knowledge of how SAS-6 regulates ciliogenesis and CD. This application is innovative because this is the first study investigating the consequences of the MCPH-associated sas-6(L69T) mutation in an in vivo model system. Hence, we expect that the successful completion of the aims of this proposal will provide knowledge that will vertically advance the field. Moreover, this proposal is significant because there is currently no cure or treatment options for MCPH. A clearer understanding of the consequences of this mutation will enable a better understanding of the mechanism of incidence and progression of MCPH.
Former Projects
Phase 1 Graduates
Hui-Ying Lim, Ph.D.
Associate Professor of Physiology, University of Oklahoma Health Sciences Center
“Reactive oxygen species in the epi/pericardium regulate Drosophila heart physiology“
Lorin Olson, Ph.D.
Associate Member, Cardiovascular Biology Program, OMRF
“Understanding connective tissue development and disease with PDGFR-driven models of fibrosis”
Roberto Pezza, Ph.D.
Associate Member, Cell Cycle and Cancer Biology Program, OMRF
“The roles of Hop2 and Mnd1 in mouse meiotic homologous recombination.”
Chris Sansam, Ph.D.
Associate Member, Cell Cycle and Cancer Biology Program, OMRF
“TopBP1 and TICRR in the chemotherapy response and embryonic development”
Weidong Wang, Ph.D.
Associate Professor of Medicine, Endocrinology & Diabetes
William K. Warren, Sr. Chair in Diabetes Studies
“Derivation of pancreatic beta cells from human induced pluripotent stem cells”
Phase 2 Graduates
David Forsthoefel, Ph.D.
Assistant Member, Genes & Human Disease, OMRF
“Non-autonomous regulation of stem cell proliferation during regeneration”
Jian Li, Ph.D.
Assistant Professor of Cell Biology & Anatomy, New York Medical College
“Spatiotemporal regulation of stress-responsive transcription factors in organismal aging”
Gaurav Varshney, Ph.D.
Assistant Member, Genes & Human Disease, OMRF
“Functional genomics of hearing loss genes in zebrafish”
Wan Hee Yoon, Ph.D.
Assistant Member, Aging & Metabolism, OMRF
“Unraveling mechanisms for neurological diseases caused by ATAD3A mutations”
Cores
Bioinformatics and Pathways Core
The goal of the Bioinformatics and Pathways Core is to aid in the experimental design, analysis, interpretation and presentation of data within the systems being studied by the COBRE investigators. Jonathan Wren, Ph.D., serves as director and is assisted by Constantin Georgescu, Ph.D., a trained statistician.
The specific aims of the core are 1) to provide complete bioinformatics analysis and biological interpretation of high throughput data, 2) to identify key genes and biomarkers involved in processes of interest and predict gene functions, phenotypes, and disease relevance, and 3) to create a sustainable core facility that can be used institution-wide. The services of the core are available free of charge to COBRE investigators.
In addition to providing conventional statistical analyses, the Bioinformatics and Pathways Core has novel software developed by Wren.
GAMMA
The Global Microarray Meta-Analysis program predicts functions of poorly annotated genes based on co-expression data and mutual information metrics. GAMMA aids in interpreting the functional significance of these genes in biological studies. Conversely, it can also identify candidate genes of interest relevant to experimental systems under study (e.g., meiosis, hematopoiesis, etc.) and screen for those where no publications exist between the gene and the system, enabling discovery of novel associations for new investigators.
IRIDESCENT
A second novel software program called IRIDESCENT, or Implicit Relationship IDEntification by Software Construction of an Entity-based Network from Text, is designed for large-scale analysis of PubMed abstracts. IRIDESCENT automates the identification and analysis of relationships within the published literature by identifying simple relationships between terms, along with a relative strength of association between them. This large network of relationships between genes, diseases, phenotypes, chemical compounds, ontological categories and FDA-approved drugs serves as a basis for analyzing lists (e.g., microarray data), and for identifying implied relationships. That is, given two things that are not related themselves, IRIDESCENT can be used to identify things they have in common. By evaluating the statistical significance of what they have in common, a measure of strength of their relatedness can be developed.
Genome Runner
Genome Runner is a tool for automating genome exploration. It performs annotation and enrichment analyses of user-provided genomic regions (SNPs, ChIP-seg binding sites, etc.) against >6000 human epigenomic features available from the UCSC genome browser. It gives a detailed annotation of each genomic region in the input data and can be used to prioritize individual genomic regions by the total number of epigenomic features they co-localize with. It also provides p-values for statistically significant co-localizations of input genomie-wide data with genome annotation features selected for the analysis. These p-values can be used to prioritize epigenomic features associated with user data.
Imaging Core
The Aim of the Imaging Core is to provide enabling technology in histology and microscopy for investigators in the Developmental Biology COBRE.
Facility Overview
The Imaging Core suite in OMRF’s research tower houses sensitive instruments in dedicated rooms built over a vibration-damping sub-floor (designed by Chad Himmel, P.E., of JEAcoustics). There is also optimized space for sample preparation, tissue sectioning and off-line computing. Sample preparation equipment includes two automated tissue processors, a paraffin embedding station, four microtomes, two cryostats, and an automatic coverslipper. Imaging Core instruments include a cryo-ultramicrotome, a high-pressure freezer, slam freezers, freeze fracture equipment, a freeze substitution system and a sputter coater, as well as
- Standard upright and inverted microscopes
- Three Zeiss LSM-based laser scanning confocal microscopes (a LSM510, LSM 710 and a LSM 880 (all with spectral unmixing capabilities and environmental chambers for live cell work). The 880 is equipped with an Airyscan detector capable of live-cell super-resolution imaging
- An Olympus FV 1000 confocal microscope with incubation for live-cell imaging
- Hitachi H-7600 transmission electron microscope fitted with a Kodak 4Kx4K 4.0 camera
- Structured illumination microscope with Colibri LED illumination
- A Zeiss Axiosscan 7 whole slide scanner. This system is capable of storing 100 slides at a time and image using brightfield, fluorescent, and polarized modalities
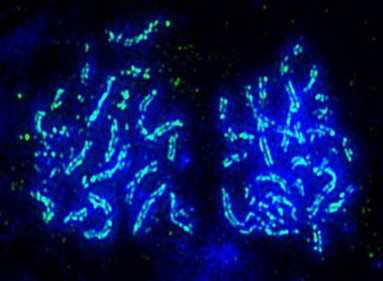
The core also houses a super-resolution DeltaVision OMX-SR microscope. As the only super-resolution microscope in Oklahoma, this instrument provides fluorescent microscopy images with much higher resolution than is possible in conventional fluorescence microscopy through two methods: structured illumination microscopy or SIM, and single molecule localization microscopy or localization microscopy. These methods effectively double or quadruple the resolution of conventional microscopy and are applicable to a wide range of research approaches. Computer support includes a 3D deconvolution 1.6TB data server and a dual zeon 500 GB data server. Also available are 3D deconvolution and data analysis workstations and extremely high-resolution printers.
The core has recently begun offering Spatial Transcriptomics services using both the Nanostring (GeoMx DSP) and 10X Genomics (Visium) platforms. These platforms allow investigators to combine histological or immunohistological information with sensitive high-plex profiling of protein or RNA transcripts at the cellular level, allowing for spatial resolution of functionally distinct cell populations and structures within the morphological context of the tissue. The systems use paraffin embedded or frozen tissue sections and fluorescent or chromogenic marker overlays to define the structural morphology of the tissue and help identify cell types or regions of interest for deeper examination. The imaging core is capable of processing the samples from tissue collection, through staining and library preparation, to sequencing.
Leadership
Gary Gorbsky PhD serves as the Director of the Imaging Core. Dr. Gorbsky is the Chair of the Cell Cycle and Cancer Biology Program at OMRF, has been an acting officer of the American Society of Cell Biology, and is an expert microscopist. He will have primary administrative and scientific oversight of the Imaging Core. Dr. Gorbsky is available to COBRE Investigators to provide advice on the proper design, implementation, and interpretation of imaging experiments.
Ben Fowler is the Imaging Core Manager and oversees all aspects to the daily operations of the Imaging Core. Mr. Fowler holds dual Master of Science degrees in Microbiology and Cell Biology. He has been managing the Imaging Core since its inception in 1999. Mr. Fowler has extensive experience in developing experimental protocols for fluorescence and electron microscopy, maintaining the full range of equipment within the Imaging Core, and in managing the personnel and business aspects of the facility.
Training
Mr. Fowler and two of the other full-time technical staff in the facility have extensive experience in training new users to utilize the full suite of light and electron microscopy equipment, following teaching protocols that have been developed over the past ten years. Access to each instrument is controlled by key-card activated doors and log-in requirements for the operating software.
Maintenance
The microscopes are covered by service contracts and Mr. Fowler performs preventative maintenance and quality control procedures as needed. Mr. Flower and his staff are able to perform common minor repairs so that down time is minimized.
Organization
Mr Fowler provides daily management of the Imaging Core while Dr. Gorbsky provides administrative oversight. Prospective users are trained in the safe and effective use of individual microscopes by Core technicians. Use of the instruments is monitored by the appropriate Core personnel until the user is approved to operate the instrument independently, including after hours.
Time allocation
User reservations are regulated by an automated, web-based calendar system which is monitored throughout normal working hours by Core personnel.












